We are excited to share a new feature in Ynet News covering the latest discovery from the Glickman Lab!
The article, “Israeli scientists reveal how brain ‘takes out the trash’, and may spread Alzheimer’s,” highlights our recent study led by Prof. Michael Glickman and postdoctoral fellow Dr. Ajay Wagh (published in PNAS).
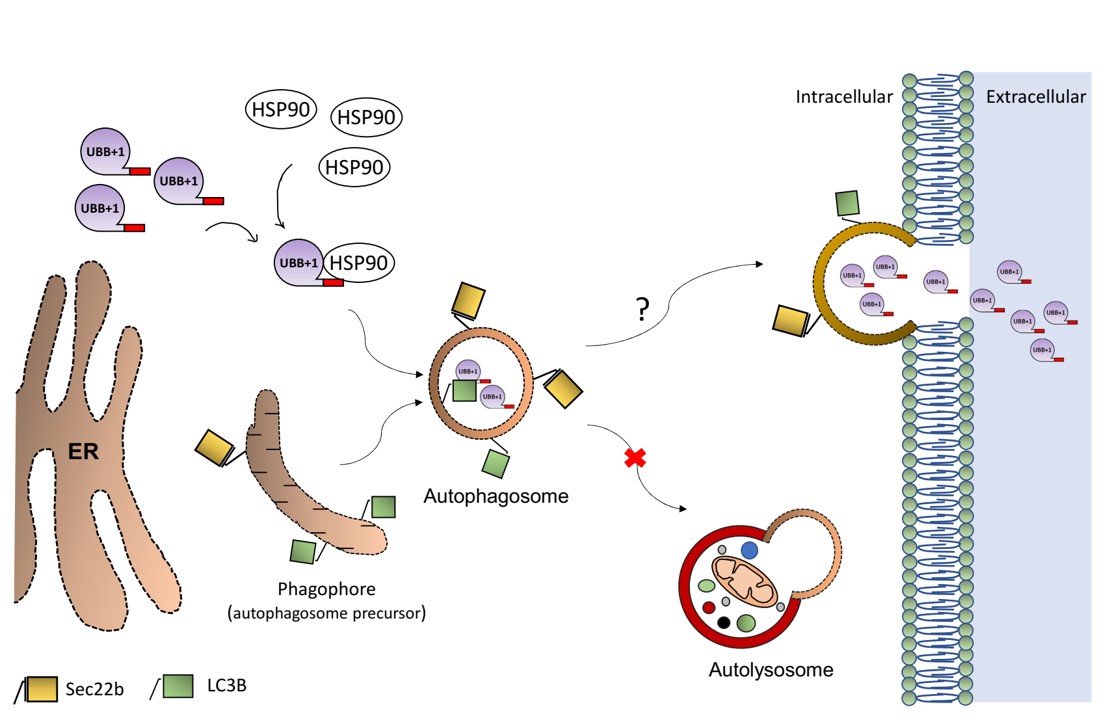
The findings: We discovered that in Alzheimer’s disease, brain cells don’t just accumulate toxic proteins—they actively expel them. While this survival mechanism saves the individual cell, it effectively “dumps the trash” on neighboring neurons, causing the disease to spread across the brain.
As Prof. Glickman explains in the interview:
“We all want someone to take out the trash… But in this case, the cells are dumping their trash on their neighbors. Although this solves an acute problem for the individual cell, it may cause long-term damage to the entire tissue.”
This insight opens new doors for early diagnosis via cerebrospinal fluid and suggests new therapeutic strategies targeting this disposal pathway.